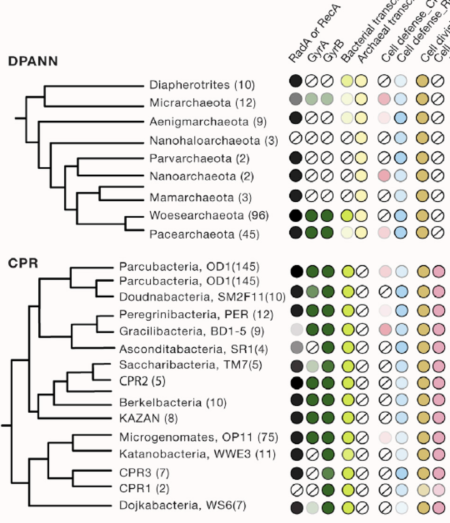
Understanding of microbial diversity has been dramatically expanded through analysis of genomes from groups of organisms previously inaccessible to laboratory-based identification and characterization.
Analysis of genomes from little-explored subsurface environments has uncovered new evolutionary patterns, including a group that may be ancestral to Eukaryotes, humanity’s own branch of life. Also evident are two major radiations of microorganisms that appear to live primarily via symbiosis with other bacteria and archaea. These organisms have ecosystem importance via impacts on their hosts, geochemical cycling, and potentially play roles in agriculture and human health.
Summary
The tree of life is arguably the most important organizing principle in biology and perhaps the most widely understood depiction of the evolutionary process. It explains how humanity is related to other organisms and where we may have come from. The tree has undergone some tremendous revolutions since the first version was sketched by Charles Darwin. A major innovation was the construction of phylogenetic trees using DNA sequence information, work that enabled the definition of the three domains of life: Bacteria, Archaea, and Eukaryotes. More recently, the three-domain topology has been questioned, and eukaryotes potentially relocated into the archaeal domain. Beyond this, and as described here, cultivation-independent genomic methods that access sequences from organisms that resist study in the laboratory have added many new lineages to the tree. Their inclusion clarifies the minority of life’s diversity represented by macroscopic, multi-celled organisms and underscores that humanity’s place in biology is dwarfed by bacteria and archaea.
Citation
C. J. Castelle and J. F. Banfield, “Major New Microbial Groups Expand Diversity and Alter our Understanding of the Tree of Life.” Cell, 172, 1181-1197 (2018) DOI: 10.1016/j.cell.2018.02.016